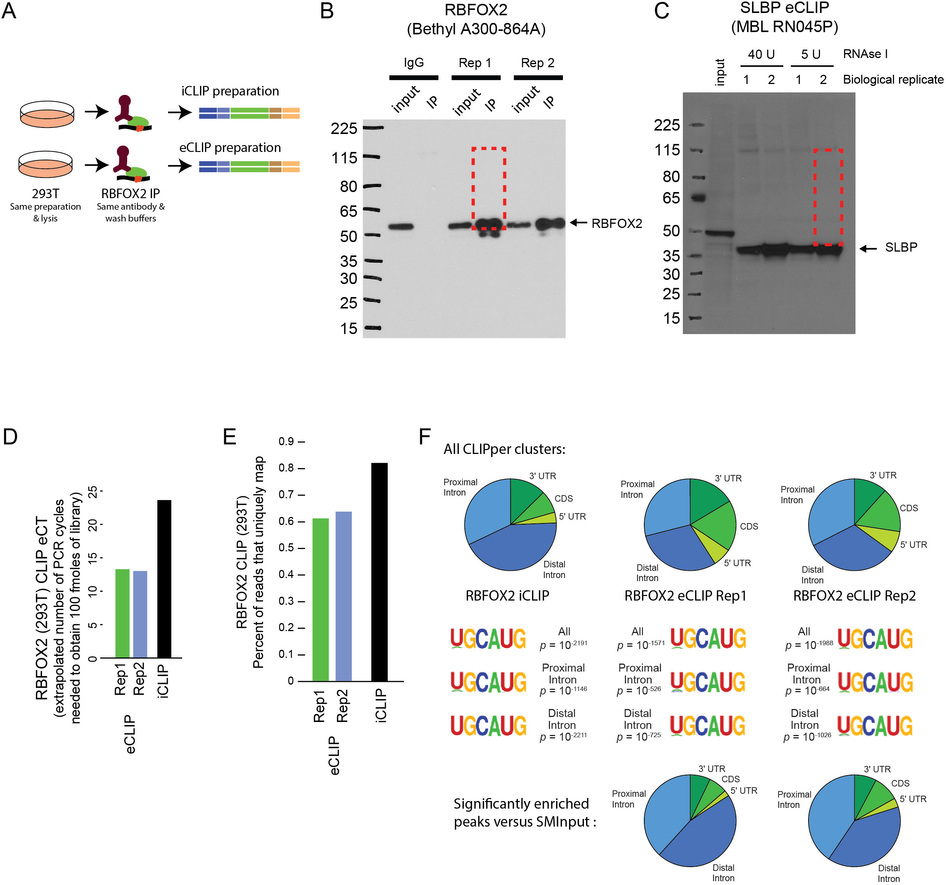
Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP) - CaltechAUTHORS
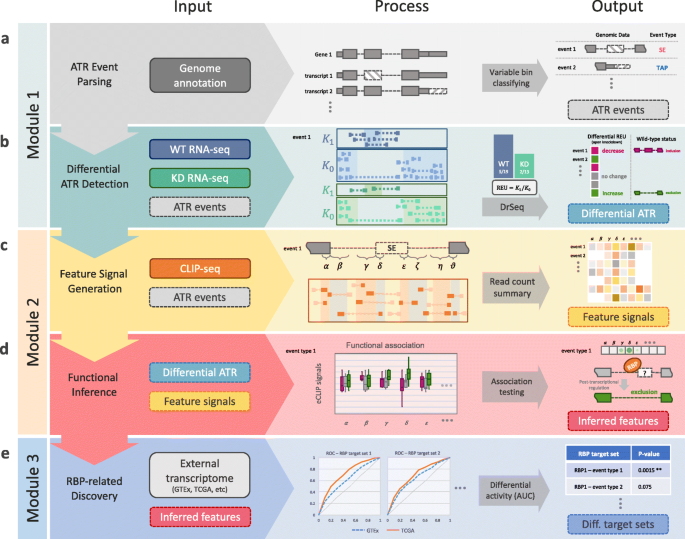
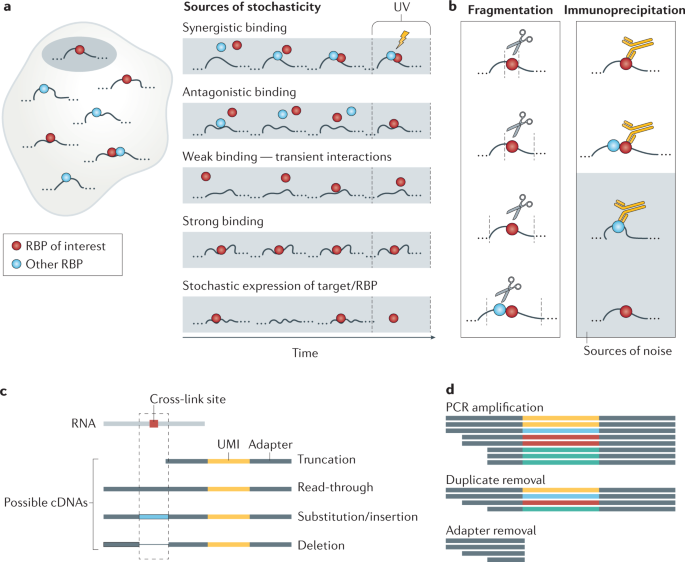
SURF: integrative analysis of a compendium of RNA-seq and CLIP-seq datasets highlights complex governing of alternative transcriptional regulation by RNA-binding proteins | Genome Biology | Full Text

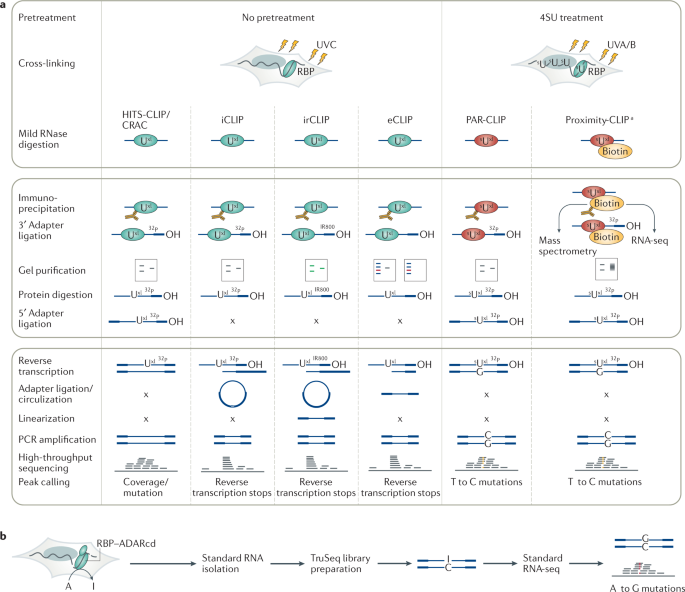
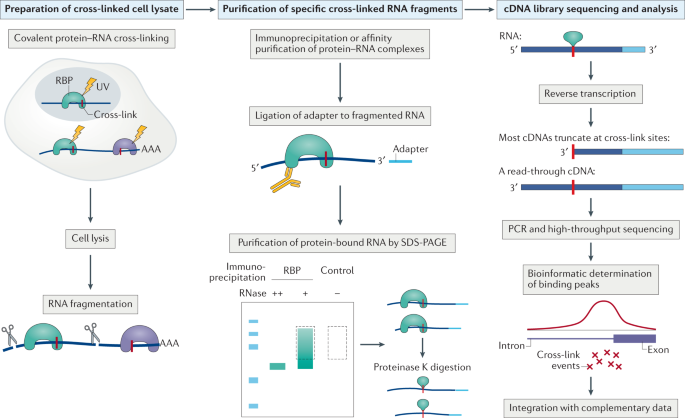
Schematic summary of CLIP and iCLIP methods. After UV cross-linking... | Download Scientific Diagram

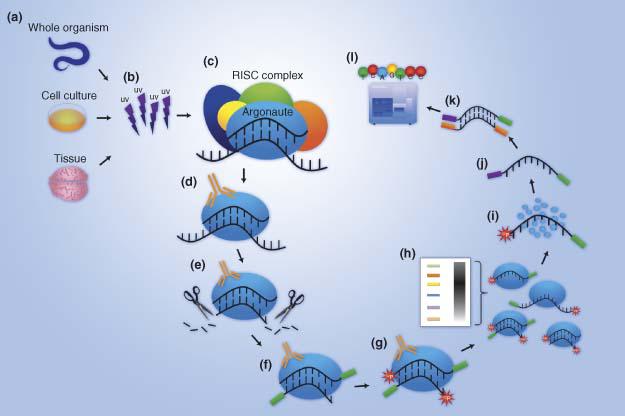
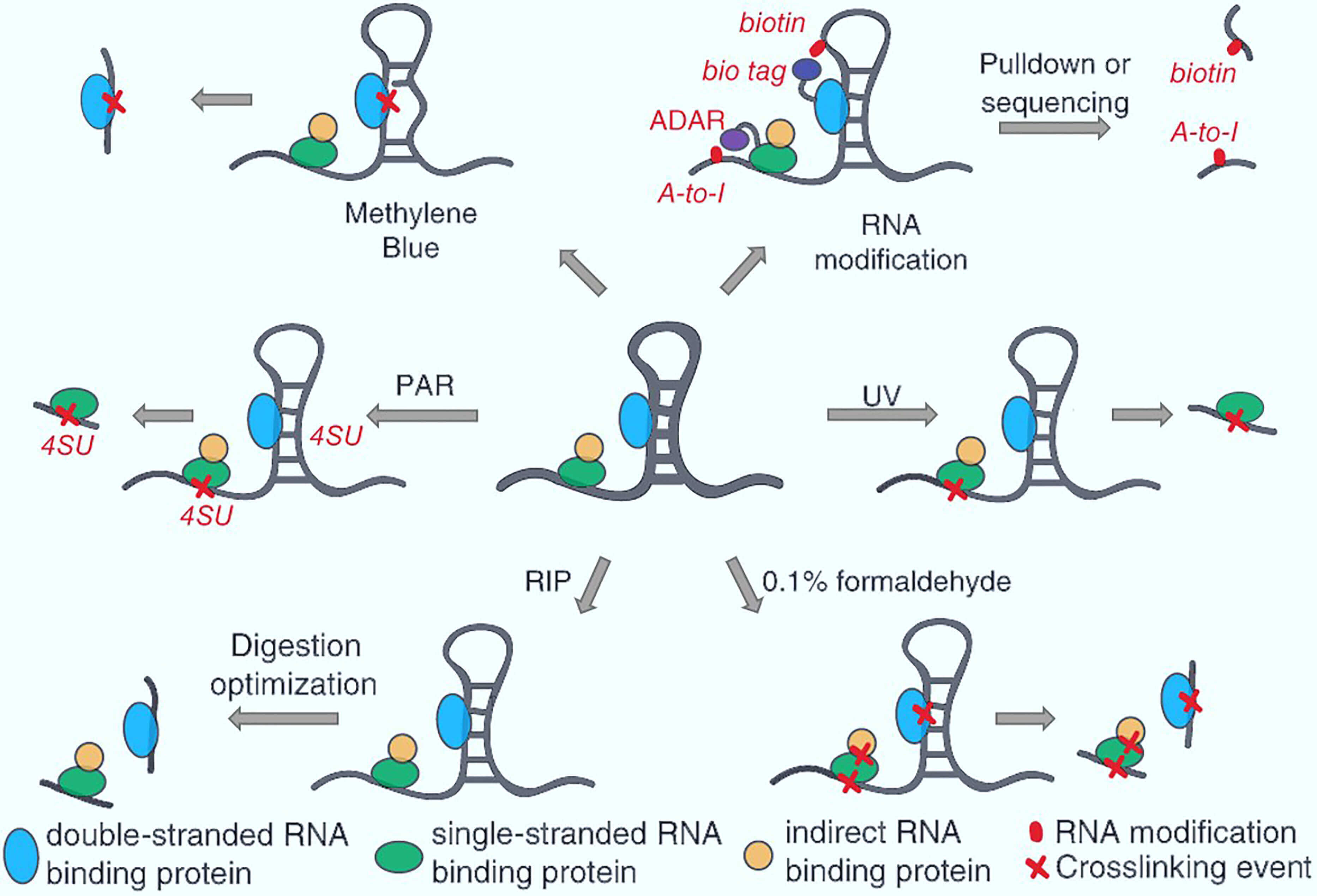
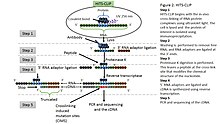
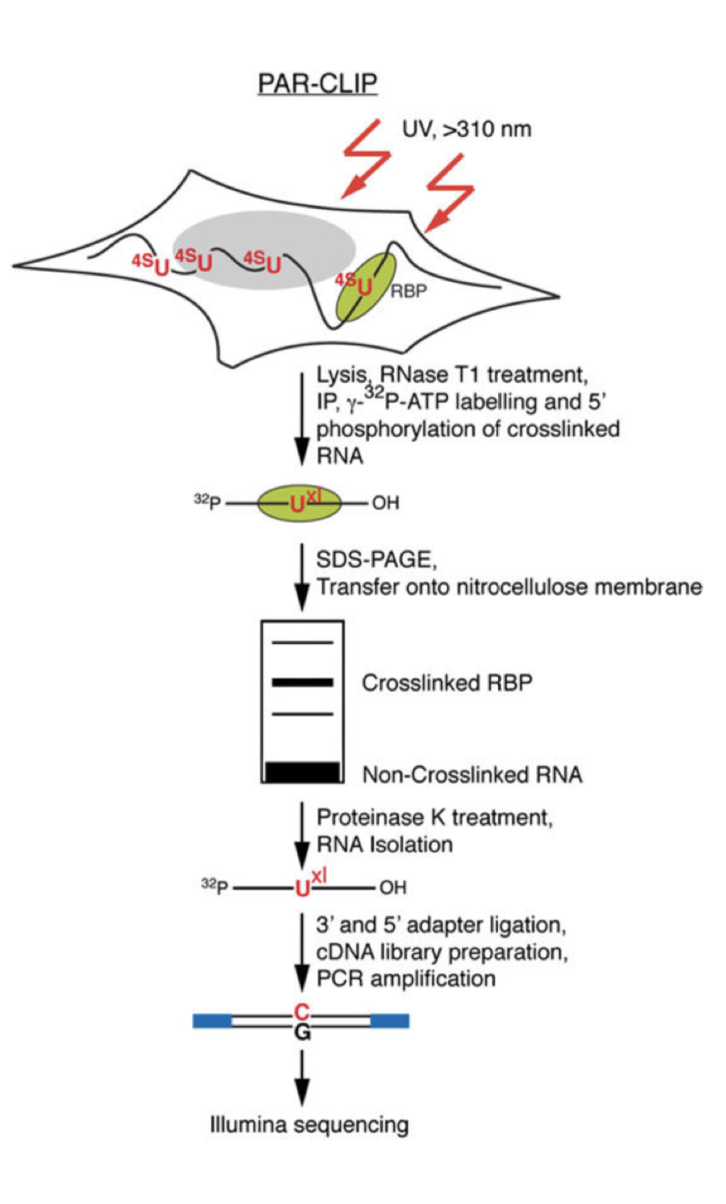
Outline of HITS-CLIP, PAR-CLIP and several variants, iCLIP, iCLAP and... | Download Scientific Diagram

Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell
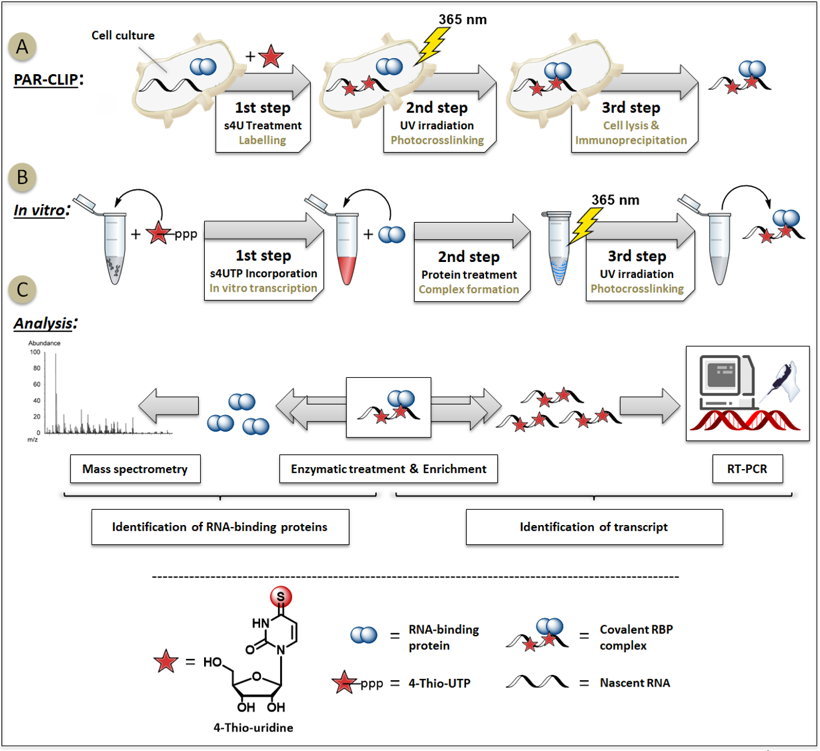
![PDF] Quantitative mass spectrometry and PAR-CLIP to identify RNA-protein interactions | Semantic Scholar PDF] Quantitative mass spectrometry and PAR-CLIP to identify RNA-protein interactions | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/85f2546f2ebc745edebbaf20459e20a394664b72/5-Figure4-1.png)
PDF] Quantitative mass spectrometry and PAR-CLIP to identify RNA-protein interactions | Semantic Scholar
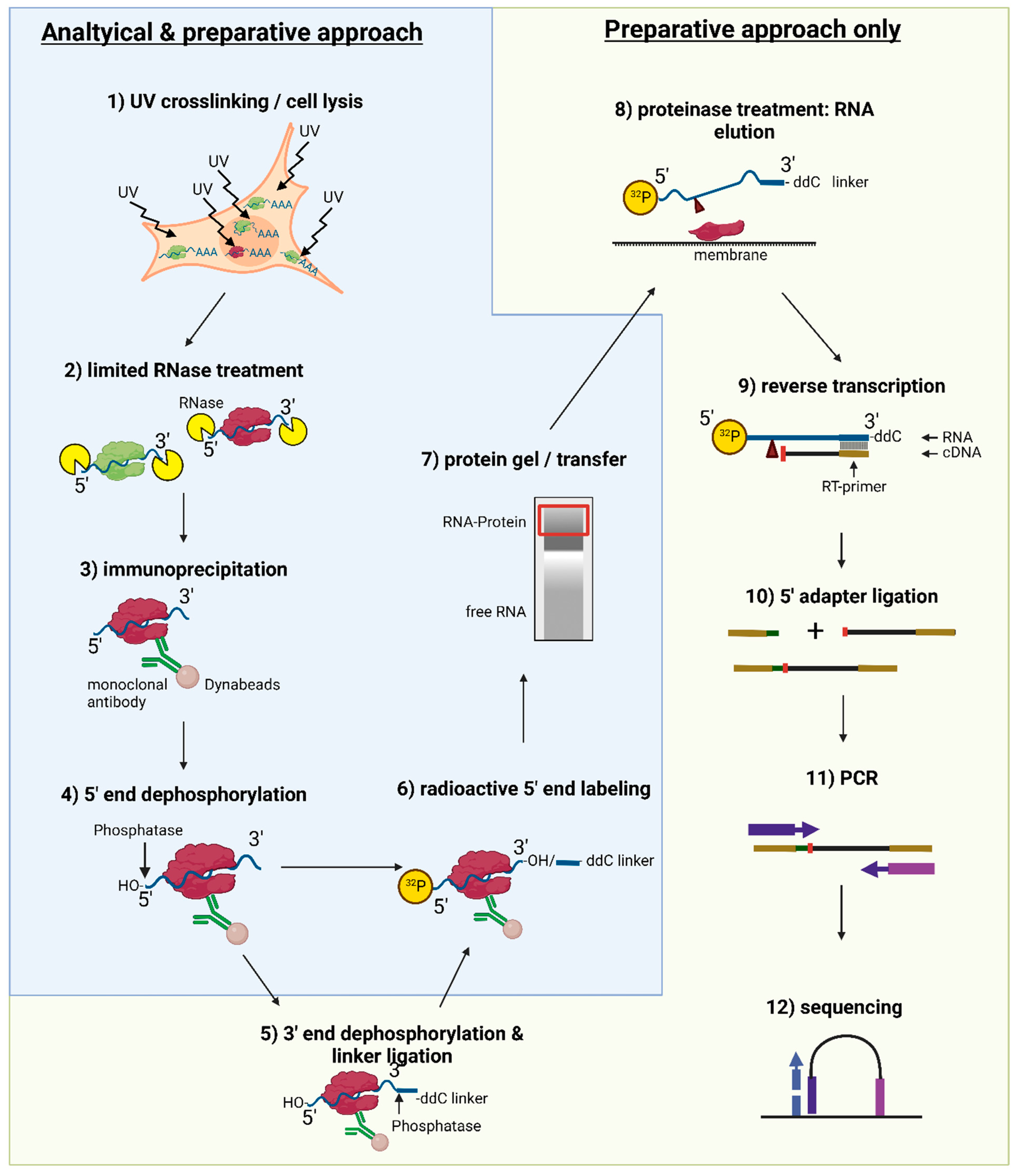
Analysis of the nucleocytoplasmic shuttling RNA-binding protein HNRNPU using optimized HITS-CLIP method | PLOS ONE
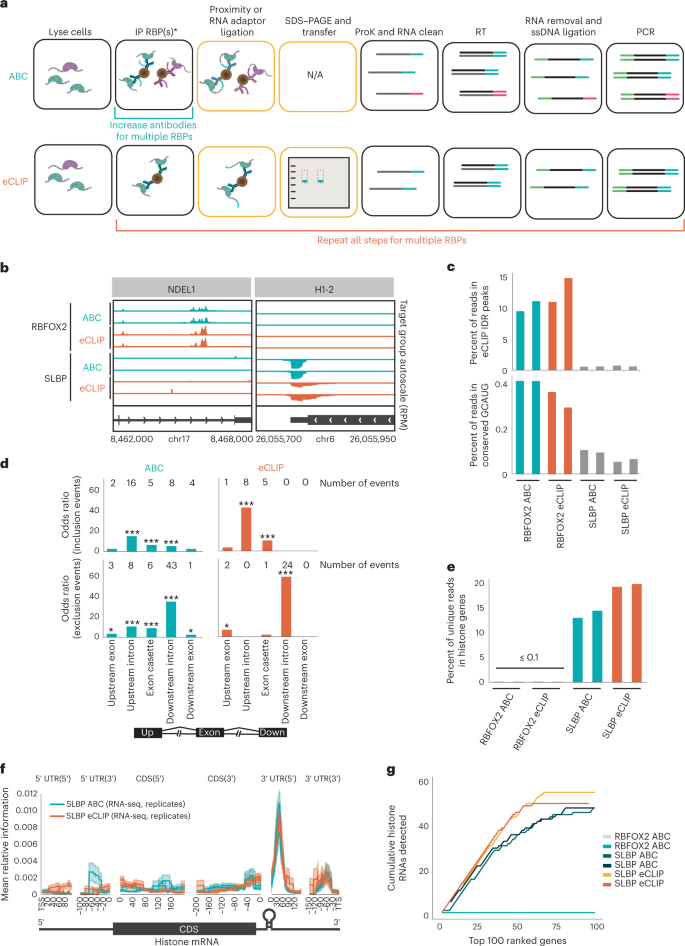
Multiplexed transcriptome discovery of RNA-binding protein binding sites by antibody-barcode eCLIP | Nature Methods
The RNA Binding Specificity of Human APOBEC3 Proteins Resembles That of HIV-1 Nucleocapsid | PLOS Pathogens



-2.jpg)